Installing GBRS
Last updated on 2024-09-06 | Edit this page
Overview
Questions
- What supporting software is needed to install GBRS?
- How do I configure my installation for my computing cluster?
- Where do I obtain the GBRS reference files?
Objectives
- Understand which supporting software is required by GBRS.
- Install supporting software and GBRS.
- Download the required reference files for Diversity Outbred mice.
Introduction
Genotyping by RNA Sequencing (GBRS) uses several pieces of supporting software to run. We use the Nextflow workflow software and Singularity containers. Your computing cluster will also have a job submission system. Examples of this are slurm or PBS.
Install Nextflow
Many computing clusters will have nextflow installed.
However, if it is not installed on your cluster, nextflow
provides
detailed
installation instructions. The installation requires a version of
Java as well.
You should contact your computing cluster administrator and ask them
if nextflow is already installed and, if so, how to access
it. Sometimes you need to add software with an explicit command
like:
module load nextflowInstall Singularity
There are several software container systems that are in popular use,
including Docker and
singularity.
Many computing clusters use Singularity because it does not
allow root access from within containers.
Again, you should contact your computing cluster administrator and
ask whether Singularity is installed and, if so, how to
access it.
Install Nextflow
Nextflow is a scripting language which runs computational pipelines. It handles running jobs and tracking progress. Most computing clusters should have this installed. If not, you will need to install it and make sure that it is in the PATH variable in your environment.
Nextflow Pipeline Repository
The Computational Sciences group at The Jackson Laboratory has
created a suite of next-generation sequencing tools using
nextflow. These are stored in a publicly available Github
repository.
Navigate to https://github.com/TheJacksonLaboratory/cs-nf-pipelines and click on the green “Code” button.
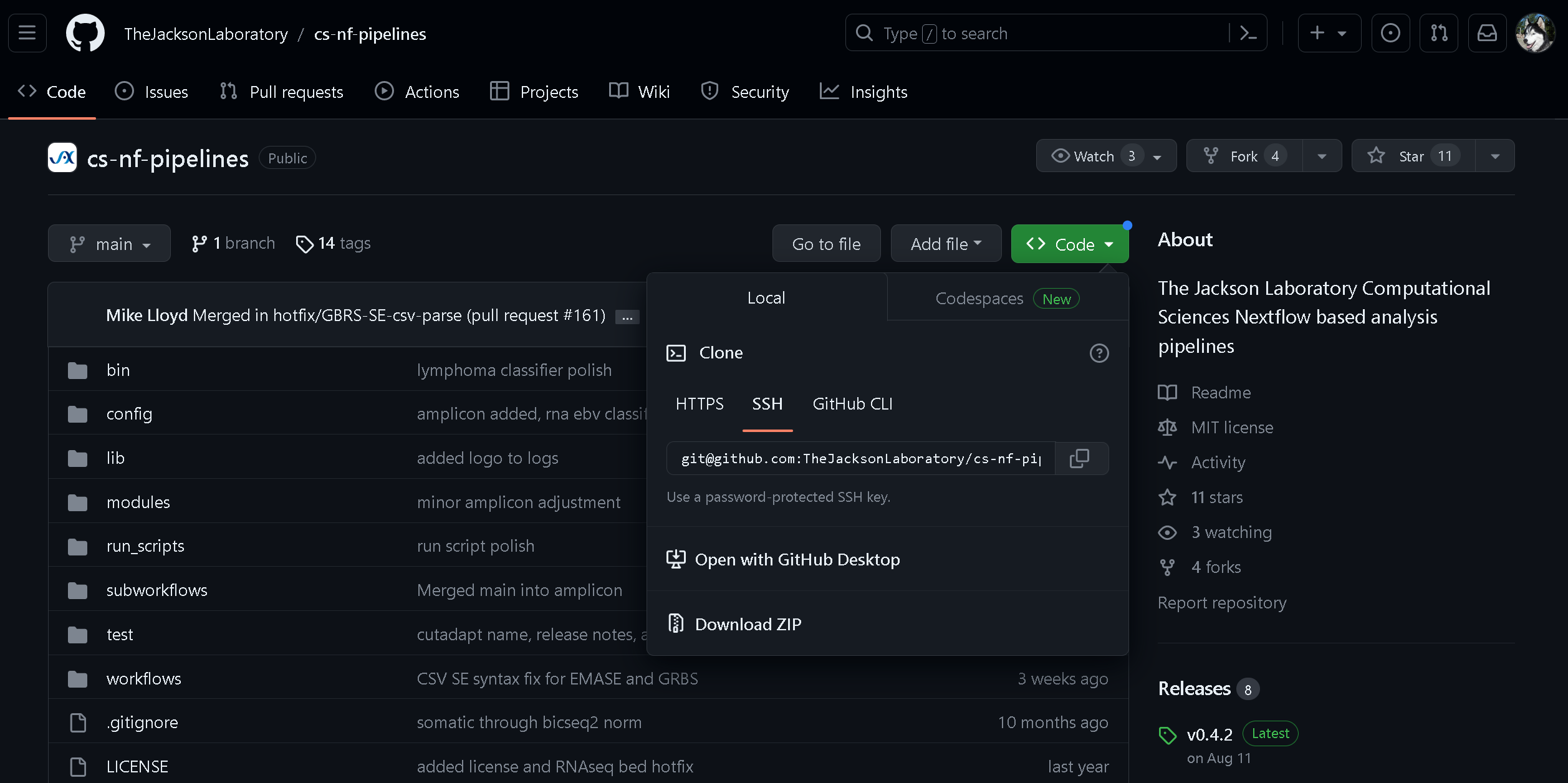
This will show a pop-over window with a code which you can copy to your clipboard. Change into a directory where you wish download the repository. Depending on how you clone Github repositories (https or ssh), your command may look something like this:
git clone https://github.com/TheJacksonLaboratory/cs-nf-pipelines.gitor this:
git clone git@github.com:TheJacksonLaboratory/cs-nf-pipelines.gitCluster Profile Configuration File
There are sample profile configuration files on Github. These files are for the clusters at The Jackson Laboratory. You will need to modify the values in this file to configure the settings for your cluster.
We provide some examples of blocks which you may need to edit.
singularity {
enabled = true
autoMounts = true
cacheDir = '/projects/omics_share/meta/containers'
}You should set the value of cacheDir to a directory in which you would like the pipeline to store software containers.
process {
executor = 'slurm'
queue = 'compute'
clusterOptions = {task.time < 72.h ? '-q batch' : '-q long'}
module = 'slurm'
}If your cluster does not use slurm, then you should edit
the ‘executor’ variable to be the one which you use. See the nextflow
executor documentation for more information.
executor {
name = 'slurm'
// The number of tasks the executor will handle in a parallel manner
queueSize = 150
submitRateLimit = '1 / 2 s'
// Determines the max rate of job submission per time unit, for example '10sec' eg. max 10 jobs per second or '1/2 s' i.e. 1 job submissions every 2 seconds.
}If your cluster does not use slurm, then you will need
to modify the block above.
Reference Data Files
The reference files for Diversity Outbred mice are stored on Zenodo. Create a directory in which you will store the reference files and change into it.
Callout
These reference files are on mouse genome build GRCm39 and use Ensembl 105 gene annotation.
You do not need all of the files. You will need the following files:
Bowtie transcript indices:
- bowtie.transcripts.1.ebwt
- bowtie.transcripts.2.ebwt
- bowtie.transcripts.3.ebwt
- bowtie.transcripts.4.ebwt
Reference genome FASTA file index:
GBRS needs the reference file index for internal bookkeeping. The reference genome is contained in the Bowtie indices above.
Gene and Transcript information:
- emase.fullTranscripts.info
- emase.gene2transcripts.tsv
- emase.pooled.fullTranscripts.info
- ref.gene_pos.ordered_ensBuild_105.npz
Marker grid on GRCm39:
GBRS uses a 74,165 pseudomarker grid with markers evenly distributed across the genome. This is the grid on which the results are reported. The motivation is that each tissue will express a different set of genes with different genomic positions and this grid allows for consistent reporting of results.
Emission probabilities:
Transition probabilities:
Colors to use when drawing founder haplotypes:
Once you have these file in place, navigate to the “Running GBRS” lesson.
Key Points
- GBRS requires support software and Diversity Outbred reference files to run.
- GBRS uses
nextflowto run the GBRS pipeline. - GBRS uses a container system, either
dockerorSingularity, to modularize required software versions. - Diversity Outbred (DO) reference files are needed to run GBRS on DO mice.
